Brain Tumor Radiomics Pipeline
Automated multimodal MRI radiomics feature extraction pipeline using 3D Slicer and NVIDIA AIAA.
In clinical and research settings, extracting quantitative features from medical images (radiomics) often requires tedious manual interaction. This case study details the development of an automated pipeline using 3D Slicer's Python scripting environment to process multimodal MRI scans (T1, T2, FLAIR, ADC) and extract radiomic features for machine learning applications.
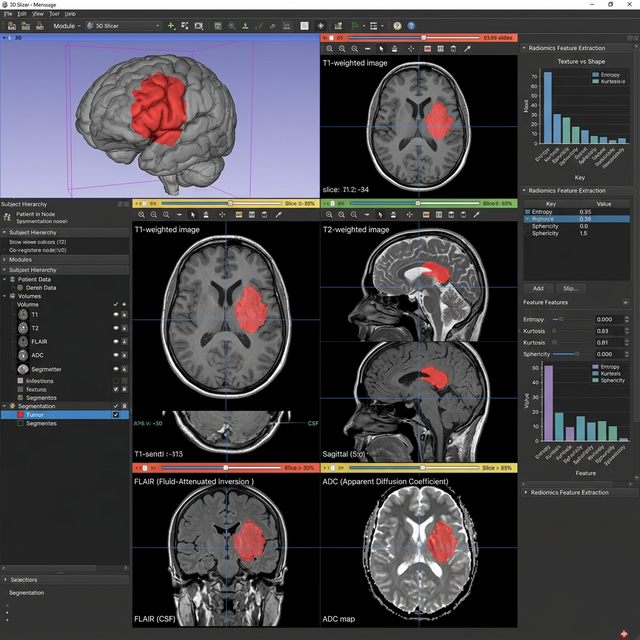
Project Overview
The pipeline is designed to eliminate the GUI button-clicking constraints of traditional Slicer workflows for high-throughput AI research. The challenge was successfully isolating relevant volumes iteratively across multiple patients, matching unique sequences using RegEx patterns, and robustly generating segmentation measurements with the SlicerRadiomics module.
- Data Parsing & Alignment: Developed a Python workflow to automatically traverse patient directories and identify T1, T2, FLAIR, and ADC volumes using regular expressions.
- AI-assisted Segmentation: Integrated NVIDIA AIAA (AI-Assisted Annotation) models via Python to create initial tumor segmentations efficiently, ensuring consistency across the dataset.
- Automated Feature Extraction: Leveraged
SlicerRadiomicsLogicusing parameter.yamlfiles to computationally extract over 100+ shape, intensity, and texture features. - Pipeline Output: Automatically exported segment statistics and feature matrices iteratively into CSV files and converted them into structured Excel files for seamless machine learning feature ingestion.
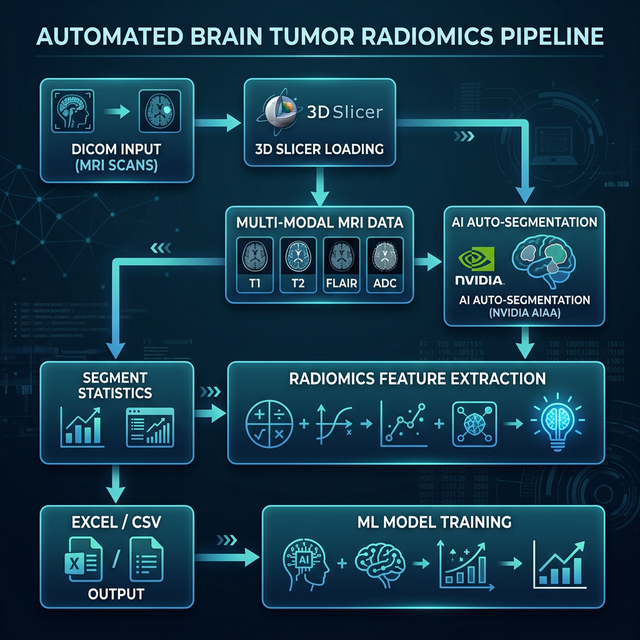
Technologies Used
- 3D Slicer 5+ (Medical Image Computing)
- Python scripting (PySide / VTK) inside Slicer kernel
- SlicerRadiomics extension
- NVIDIA AIAA (AI-assisted annotation)
- Pandas / OpenPyXL (Data structuring and manipulation)