Automated Skull Diploe Thickness Mapping
Advanced morphological tracking and segmentation of skull layers utilizing native ITK thickness filters.
Understanding variations in skull thickness, particularly resolving the complex diploe internal spongy bone layer, requires delicate segmentation. This project aimed to create a fully computational thickness map over the entire skull using programmatic control within Slicer, allowing high-throughput calculations across patient cohorts.
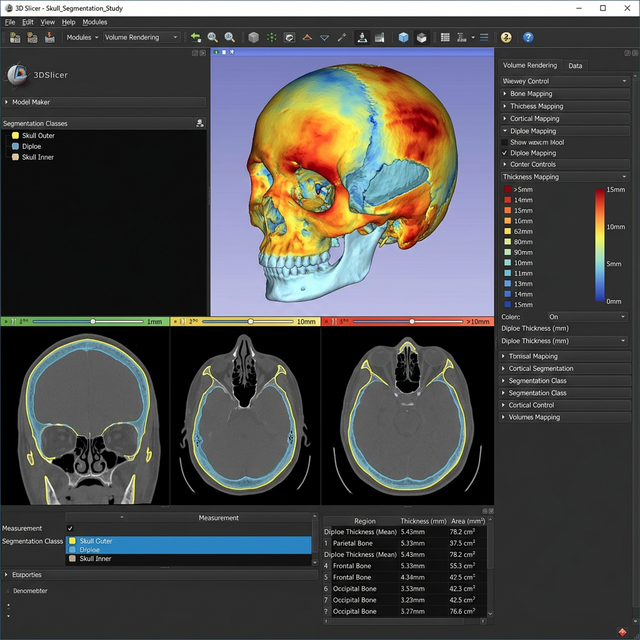
Technical Approach
The Python script fully encapsulates the complex multi-step pipeline inside Slicer. To ensure that only true skull regions were selected, automated operations parsed multiple algorithms sequentially:
- Automated Thresholding: Instantiated a class-based pipeline to programmatically connect volume nodes and invoke segment editor operations (Thresholding using specific bone density scales).
- Morphological Healing: Utilized kernel-based
operations—
KEEP_LARGEST_ISLANDandMORPHOLOGICAL_CLOSING—to remove artifacts, patch holes seamlessly, and construct the outer skull constraint shell. - Constraint-based Boolean Operations: Cloned segmentations, altered mask modes, and utilized logical overlap settings to calculate the internal diploe boundaries.
- Advanced ITK Filtering: Pulled VTK volumes to SimpleITK to execute complex
computational geometry. First,
BinaryThinningImageFilterresolved the medial axis of the bone segment. Second,DanielssonDistanceMapImageFiltercalculated the exact mathematical thickness mapping onto the object. - Data Export: Final scene reconstructions generated dynamic models containing quantitative arrays, which then systematically batch exported VTK models and color tables mapped for all folder inputs.
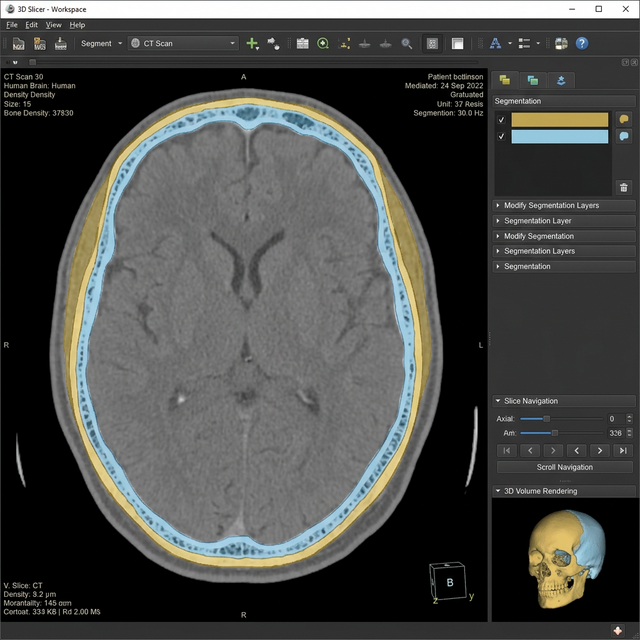
Technologies Used
- 3D Slicer 5.x
- Python Scripting
- SimpleITK & vtkITK APIs
- VTK Object Modeling